Make Python run faster
Python has a great ecosystem for machine learning, but deep learning is computationally intensive and Python is slow. In this post, I will discuss different ways that helped to make my code run faster, more specifically in physics simulation and reinforcement learning for character animations. Nevertheless, most of the tips are applicable to all computationally intensive programs.
With that, here are the 7 ways to make any Python code run faster:
- Make Your Machine Run Faster
- Try Different Python Versions and Distributions
- Profile and Optimize
- Be Mindful of Type Conversions
- Be Strategic About Memory Allocations
- Be Clever When Writing If-Statements
- Be Cautious When Using Packages
About me — I am a Ph.D. student at the University of British Columbia, and my research uses deep learning and reinforcement learning to solve character animation problems. For example, check out this endlessly walking robot and interactive web-based character controller. One of the most frustrating experiences working with reinforcement learning is how slow it can be. Even though neural network models in animation and robotics are relatively small, an experiment can still take hours and days due to the nature of reinforcement learning. The examples I’m sharing today to make Python faster are based in NumPy and PyTorch, so some experience with them will be helpful.
1 Make Your Machine Run Faster
There are only two ways to make any program run faster — write more efficient code or make your machine run faster.
Before spending hours optimizing your code, it is worth spending a few seconds to check if your machine is running as efficiently as possible. It may sound obvious, but that is precisely why it can be so easily overlooked, even by most of the front page results on Google.
Computers don’t run at maximum capacity by default. Running at peak capacity wastes energy for little to no observable benefit for daily tasks like checking emails. So the default power setting is usually on demand or balanced. This allows the CPU to use more power when necessary, such as when opening a heavy application, while saving energy when the machine is idle.
The operating system intelligently decides when the CPU can draw more power. But when we are after the maximum performance, it is better to make the decision ourselves. Running CPU in performance mode made a 10-25% difference for my codebase since most physics simulations are CPU-bound. I use the following commands to set the CPU performance mode on Linux.
sudo apt install cpufrequtils
# Check current governor
cat /sys/devices/system/cpu/cpu*/cpufreq/scaling_governor
# See options: conservative, ondemand, powersave, performance
cpufreq-info
# Turn on `performance` mode for all cores
# $(($(nproc)-1)) returns the number of cores minus 1
sudo bash -c 'for i in $(seq 0 $(($(nproc)-1))); \
do cpufreq-set -c $i -g performance; done'
# Revert back to `ondemand` mode for all cores
sudo bash -c 'for i in $(seq 0 $(($(nproc)-1))); \
do cpufreq-set -c $i -g ondemand; done'If you are running Windows, it is probably best to switch to Linux. The last time I compared running experiments on both operating systems, Linux was slightly faster. If things have changed, do let me know; I would happily switch to whatever is fastest now.
2 Try Different Python Versions and Distributions
Python, like other software, is still improving over time. Every release of Python comes with different optimizations; for example, here are the release note for Python 3.10 and Python 3.11. Newer Python versions are not necessarily faster than older versions. Try them all out and see which version works best for your codebase. Fortunately, tools like miniconda and pyenv make managing different Python versions easy.
Don’t stop at trying different Python versions, also try out different Python distributions. I found precompiled Python from conda-forge to be fast.
conda config --add channels conda-forge
conda config --set channel_priority strict
conda create -n 3.9.5 python=3.9.5 -c conda-forgePyston improves the training time by 10% on our brachiation project. I used PystonConda 2.3.4 and it was relatively painless to install all the libraries I needed, e.g., PyTorch and PyBullet. Pyston-lite is a lightweight alternative from the developers of Pyston, but I did not see significant improvement using it. Intel Distribution for Python could be an alternative to look into, but it was not faster for me. I have not tried Cinder or Python 3.11+.
The time investment in finding the best Python version and distribution is worth it if you are running reinforcement learning experiments 12 hours at a time. At a modest 5% speedup, it will save just over one hour every day.
3 Profile and Optimize
Most tips for speeding up Python code from Google’s first page results are worthwhile — follow them when possible. Most importantly, profile your code, find the bottleneck, and then optimize.
For profiling, there are many tools like py-spy and scalene. I personally cannot remember the input flags, so I tend to use the built-in profiler.
pip install snakeviz
python -m cProfile -o temp.dat _command_
snakeviz temp.datThis is what the summary of the result looks like. In this example, the list comprehension at line 6 of test.py took 14.5 seconds and the total run-time was 17.8 seconds. It’s easy to see at a glance which part of code takes the longest to execute, and therefore should be optimized if possible.
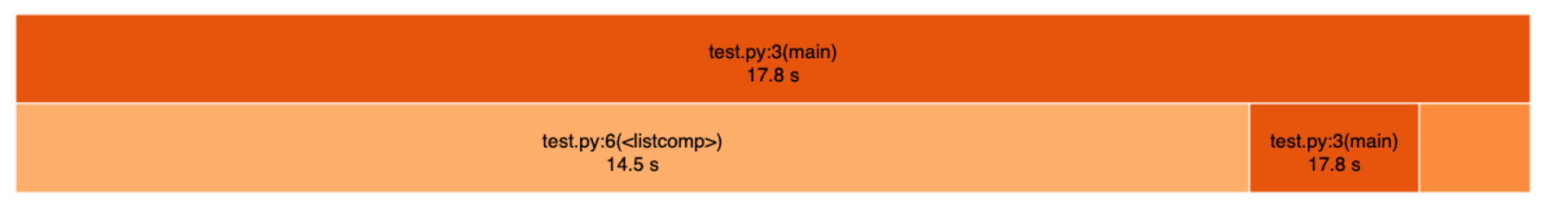

After finding the bottleneck, it is usually pretty obvious how the code can be improved. There are many resources one Google search away, so I won’t duplicate them all here. The most helpful tips for me were: use built-in functions, prefer list comprehension over loops, vectorize code using numpy, numpy can be slower for smaller arrays, check out numba and bottleneck.
Most reinforcement learning code follows a similar format, like this one:
for _ in range(epochs):
for _ in range(steps_per_epoch):
run_simulation_step()
store_training_data()
for _ in range(training_epochs):
calculate_loss()
update_model()Usually, simulation is the bottleneck because it runs on a CPU (exceptions: Isaac Gym and Brax), while everything else can be parallelized on a GPU. For perspective, a typical experiment in physics-based 3D character animation runs on the order of millions and sometimes billions of simulation steps. For training image or language models, the computation is comparable or even more sizable.
At this magnitude, even the most trivial improvement can make a big difference in the code execution time. As the ex-director of AI at Tesla recently found out, a simple one-line change swapping out np.sqrt for math.sqrt sped up data loader by 10%.
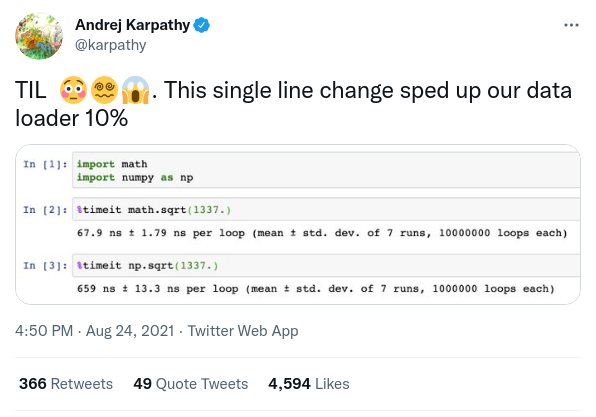
Unfortunately, there are too many tricks like this in Python. It won’t be productive to enumerate them all here, nor is it worthwhile to memorize them. The most universal strategy for writing faster code is to profile and optimize. The remaining sections describe some of the common places where more performance can be squeezed.
4 Be Mindful of Type Conversions
Python packages like numpy and torch are very convenient to use, so much so that we sometimes use them without thinking about the underlying overhead. How often have we used a numpy function just because we are too lazy to check whether the underlying object is a float, list, or numpy array?
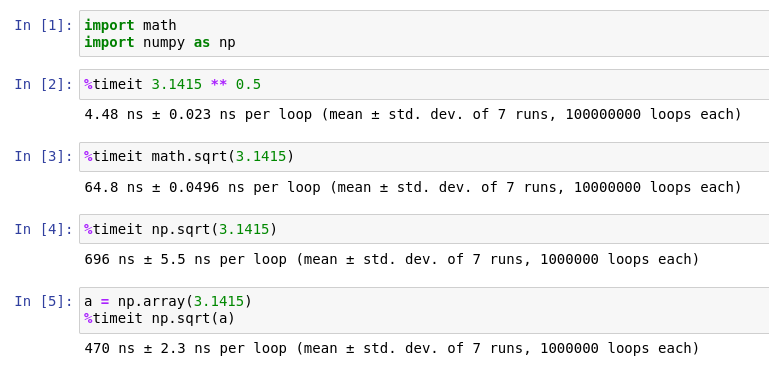
For these packages to be convenient, it must check the data type under the hood, or convert the object into something it understands. This takes time, and the nanoseconds quickly add up in a large application, especially in machine learning. In this example, manually converting the constant from float to a numpy array makes the operation 30% faster. And the fastest implementation uses the built-in power operator. Of course, the implementations behave differently when complex numbers are needed.
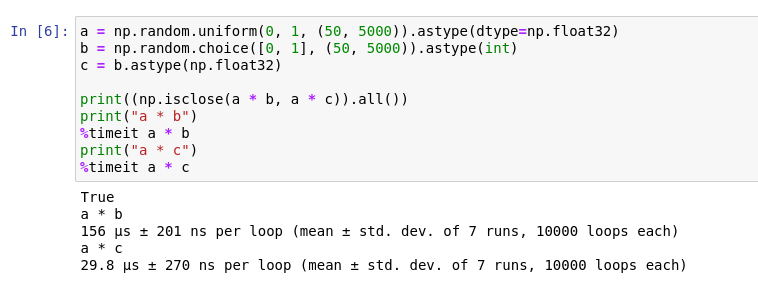
Here is another example. In machine learning, it is common to write code that uses one array to mask out another, e.g., image segmentation. There is usually a penalty for combining different data types in the same operation, and it is easy to overlook.
Being mindful about type conversions also forces us to think about the type of the underlying data object. Doing so in our brains reduces the work that needs to be done with the code. Ultimately, executing less code is faster than executing more code.
5 Be Strategic About Memory Allocations
Other than Visual Basic, the first language I learned was C in my first year of undergrad. After learning about malloc and free in class, one of the first things I learned is that allocating memory is much slower than basic arithmetic operations. Python doesn’t have a malloc function, but poor memory management can still slow down our code.
After spending a lot of time profiling and optimizing my RL simulation environment, I eventually reached a point where np.array is taking the most time in my code. At that moment, I realized that every np.array has an underlying malloc for allocating the necessary memory to store the array. And whenever I convert a list to an array, often out of laziness, I am forgetting about one of the most important lessons I learned in C.
I use PyBullet to simulate character movements for most of my projects. It is a Python binding on top of the physics engine written in C++. By default, the PyBullet API returns objects as Python tuples, e.g., robot joint angles, character root positions, etc. I will then convert these tuples to numpy arrays in Python for further computations, such as calculating the distance between two points.

Although creating a numpy array is fast, the nanoseconds accumulate and become a significant portion of the overall time. It is generally better to allocate all the memory a piece of code will need at the beginning. In reinforcement learning, this translates to allocating the replay buffer ahead of time, and directly modifying the buffer data in the environment, instead of creating new tuples every time. Similarly, in-place operations in PyTorch should be preferred over their out-of-place counterparts.

One more note about memory allocation: CPU and GPU memory should be treated separately. Transferring data between CPU memory and GPU memory takes time, i.e., torch.randn(10).to(device). For heavy computation tasks and embarrassingly parallelizable problems, it could be worthwhile to leverage GPU acceleration. But if the implementation requires transferring data back-and-forth between CPU and GPU, then any speedup from the GPU could be easily offsetted by the memory transfer time.
6 Be Clever When Using If-Statements
Remember I had previously said that executing less code is faster than executing more code? Well, it isn’t always true — sometimes mode code is faster.
The example below compares two clip_grad_norm implementations. It is pretty popular, and you might have already seen it in many open-source machine learning projects. The first implementation (with if-statement) is slightly modified from PyTorch’s source code, keeping only the necessary lines for clipping. The second implementation replaces the if-statement with a clamp operation.
The clamping technique is common when vectorizing code in numpy. In short, the second implementation clamps the multiplying factor to one and then always executes the loop. On the other hand, the first implementation checks if the multiplying factor is less than one before executing the loop. The if-statement implementation executes less code and so theoretically should be faster?

To make sure the comparison is fair, I initialize the same model every time. Despite executing more code, the second implementation is faster by 15%. If you are not convinced, here is a simple setup comparing directly the difference between two implementations using if and clamp.
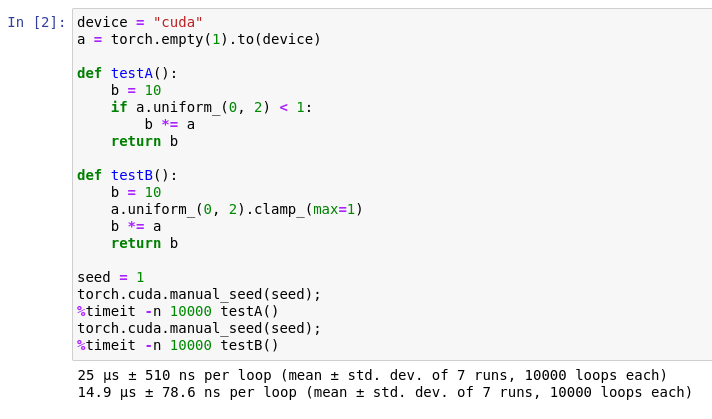
The reason why if-statements can hurt performance is because of instruction pipelining. If-statements and other branching instructions are slow because the processor doesn’t know what to execute next until the if-condition is evaluated — known as branching hazards. Normally, this isn’t a problem because the branch predictor can predict the outcome ahead of time with high accuracy. But in our second example, the randomness of the if-condition makes the branch predictor ineffective. There isn’t a one-size-fits-all solution. Try both ways and see which one is faster.
7 Be Cautious When Using Packages
The Python ecosystem is great for rapid prototyping because it has a large user base and many packages are readily available. For machine learning, other people have taken the time to write fast neural network implementations that are probably orders of magnitude faster than our own, e.g., PyTorch, TensorFlow, and Hugging Face.
But it is not always the case that off-the-shelf code will be faster than our own implementation. First, packages with a large user base need to be user-friendly and maintain backward compatibility. There can be a trade-off between performance and supporting the general use case. Second, open-source software is supported by its users who may not be too different from you or me. It simply takes too much time for individuals to find the most optimal implementation at all times, especially if they are making contributions in their free time.
One particular example is PyTorch’s utility for creating mini-batches: BatchSampler and SubsetRandomSampler. I found my own implementation for creating mini-batches is 10x faster than using the provided utility, at least in my experiments.
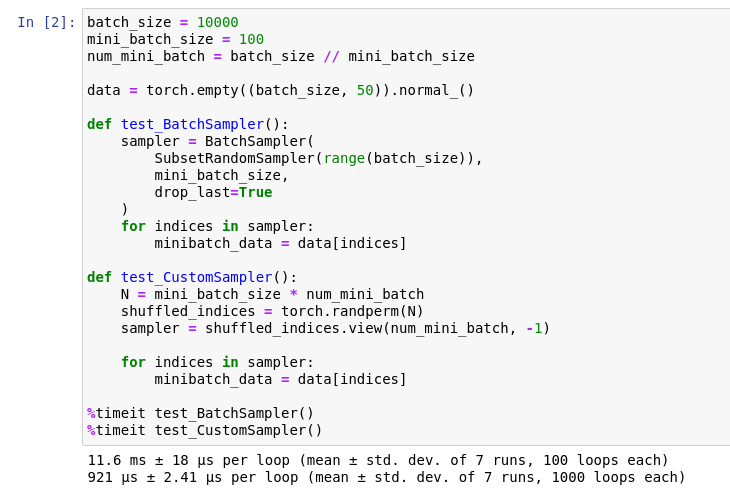
Although both implementations have approximately the same number of characters to type, the custom sampler is faster and, subjectively, easier to understand. I am sure there are some edge cases I haven’t considered, but I haven’t encountered any problem since switching to use my own implementation in my projects. For my own research projects, I prefer fast but specific code over slow but general code.
Summary
I hope this post showed you how to write faster Python code beyond swapping out loops for list comprehensions. Now I’d like to hear what you have to say. Which technique from this post is new to you? What is another trick that every programmer should know?